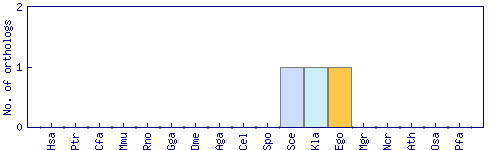
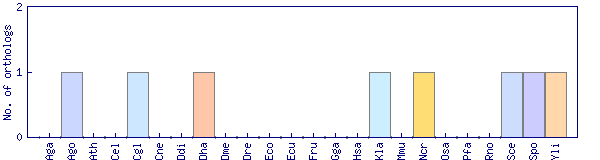
| Species | Saccharomyces cerevisiae (Sce) | ||||||||||
| Primary name | DEP1 | ||||||||||
| SGD link | S000000011 | ||||||||||
| Alternative ID | YAL013W | ||||||||||
| Description | Transcriptional modulator involved in regulation of structural phospholipid biosynthesis genes and metabolically unrelated genes, as well as maintenance of telomeres, mating efficiency, and sporulation | ||||||||||
| Synonyms | YAL013W | ||||||||||
| Datasets |
|