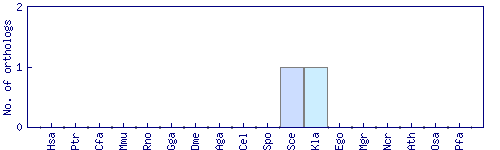
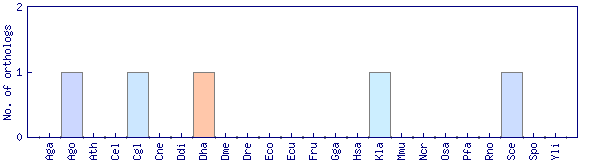
| Species | Saccharomyces cerevisiae (Sce) | ||||||||||
| Primary name | MMS4 | ||||||||||
| SGD link | S000000302 | ||||||||||
| Alternative ID | YBR098W | ||||||||||
| Description | Subunit of the structure-specific Mms4p-Mus81p endonuclease that cleaves branched DNA; involved in recombination, DNA repair, and joint molecule formation/resolution during meiotic recombination | ||||||||||
| Synonyms | YBR098W | ||||||||||
| Datasets |
|