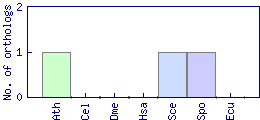
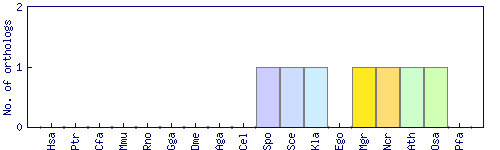
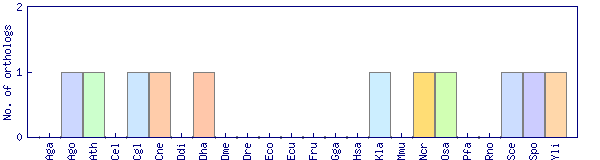
| Species | Saccharomyces cerevisiae (Sce) | ||||||||||
| Primary name | HIS6 | ||||||||||
| SGD link | S000001282 | ||||||||||
| Alternative ID | YIL020C | ||||||||||
| Description | Phosphoribosyl-5-amino-1-phosphoribosyl-4-imidazolecarboxiamide isomerase, catalyzes the fourth step in histidine biosynthesis; mutations cause histidine auxotrophy and sensitivity to Cu, Co, and Ni salts | ||||||||||
| Synonyms | YIL020C | ||||||||||
| Datasets |
|