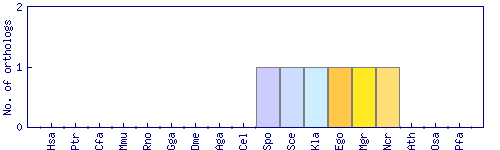
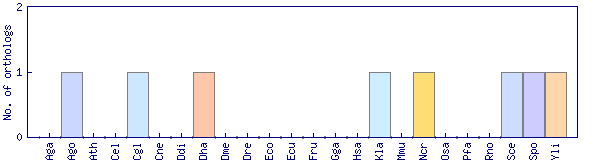
| Species | Saccharomyces cerevisiae (Sce) | ||||||||||
| Primary name | HIR3 | ||||||||||
| SGD link | S000003901 | ||||||||||
| Alternative ID | YJR140C | ||||||||||
| Description | Subunit of the HIR complex, a nucleosome assembly complex involved in regulation of histone gene transcription; involved in position-dependent gene silencing and nucleosome reassembly | ||||||||||
| Synonyms | YJR140C | ||||||||||
| Datasets |
|