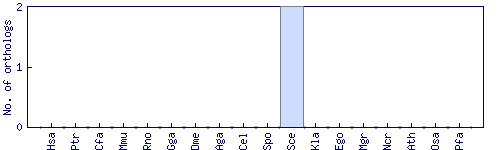
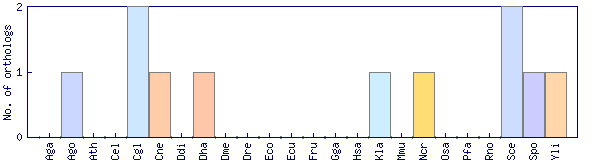
| Species | Saccharomyces cerevisiae (Sce) | ||||||||||
| Primary name | ZDS1 | ||||||||||
| SGD link | S000004886 | ||||||||||
| Alternative ID | YMR273C | ||||||||||
| Description | Protein that interacts with silencing proteins at the telomere, involved in transcriptional silencing; has a role in localization of PKA subunit Bcy1p; implicated in mRNA nuclear export; involved in mitotic exit through regulation of Cdc14p | ||||||||||
| Synonyms | YMR273C | ||||||||||
| Datasets |
|