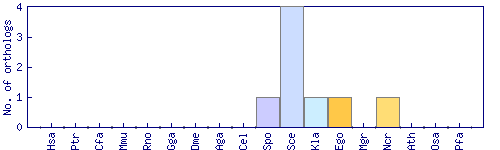
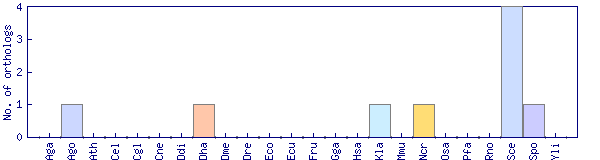
| Species | Saccharomyces cerevisiae (Sce) | ||||||||||
| Primary name | THI12 | ||||||||||
| SGD link | S000005276 | ||||||||||
| Alternative ID | YNL332W | ||||||||||
| Description | Protein involved in synthesis of the thiamine precursor hydroxymethylpyrimidine (HMP); member of a subtelomeric gene family including THI5, THI11, THI12, and THI13 | ||||||||||
| Synonyms | YNL332W | ||||||||||
| Datasets |
|